Leaving Community
Are you sure you want to leave this community? Leaving the community will revoke any permissions you have been granted in this community.
 Signaling Pathways Project (SPP): A multi-omics knowledgebase for cellular signaling pathways
Signaling Pathways Project (SPP): A multi-omics knowledgebase for cellular signaling pathways
WHAT IS IT?
Signaling Pathways Project (SPP) is a FAIR data knowledgebase that allows researchers to ask questions across thousands of annotated transcriptomic and ChIP-Seq datasets. Its powerful consensome meta-analysis pipeline allows researchers to make high-confidence connections between transcriptonal targets, their upstream regulatory pathways, and disease states.
WHO CAN USE IT?
Any researcher who is interested in identifying transcriptional targets for receptors, enzymes and transcription factors. SPP is designed specifically for bench scientists and no computer programming skills are required to use it.
WHAT TYPE OF QUESTIONS CAN IT ANSWER?
1. What cell cycle genes are regulated by FGF receptors in human liver?
2. What human genes are most responsive to insulin receptor signaling?
3. What genes in my gene set are regulated by E3 ubiquitin ligases?
STORIES OF DISCOVERY
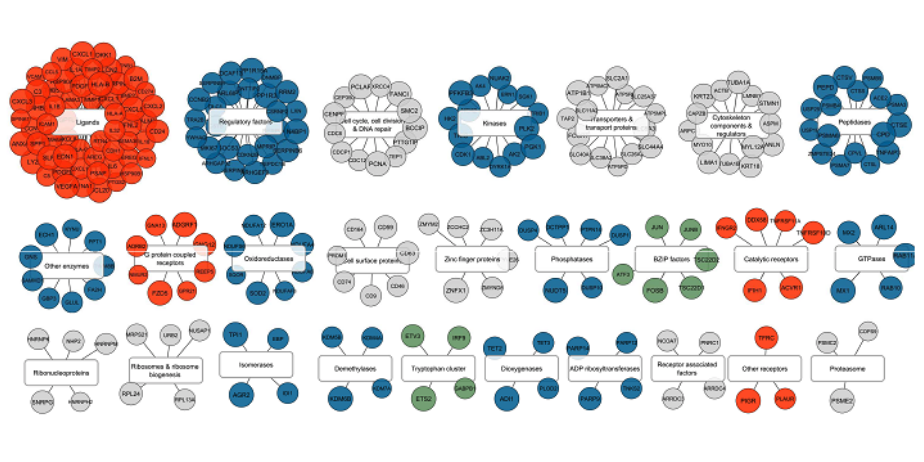
The Signaling Pathways Project Brings New Insight into COVID-19 Research
Based on evidence found using SPP, researchers hypothesized that the suppression of the interferon response to SARS2 infection by elevated...[more]
SEARCH FOR A LIST OF DATASETS THAT SPP INCORPORATES
SEARCH FOR GENES WITH IMPORTANT ROLES IN DK-RELEVANT RECEPTORS, ENZYMES, ORGANS AND TISSUES USING OUR *CONSENSOME APPROACH:
Transcriptomic Consensomes examples:
- Insulin receptor
- PPAR family in the mouse metabolic system
- Cyclin-dependent kinases in the human female reproductive system
- Mouse adipose tissue
- Mouse liver
Chip-Seq Consensomes examples
*Consensomes are list of genes ranked according to a meta-analysis of their differential expression in publicly archived transcriptomic datasets involving perturbations of a specific signaling pathway in a given biosample category. Consensome are intended as a guide to identifying those genes most consistently impacted by a given pathway in a given tissue context.
DEFINE NOVEL DK-RELEVANT SIGNALING PATHWAYS USING REGULATION REPORTS
Search by single gene examples
- CD44 gene (transcriptomic, ChIP-Seq)
- PDK4 (transcriptomic, ChIP-Seq)
Search by Gene Ontology terms examples
- Regulation of fatty acid beta-oxidation by catalytic receptors (transcriptomic)
- Regulation of inflammatory response genes by acetyltransferases (transcriptomic)
- Regulation of glycolytic process by G-protein coupled receptors (transcriptomic)
- Promoters of carbohydrate biosynthesis enzyme-encoding genes bound by E3 ubiquitin ligases (ChIP-Seq)
- Promoters of gluconeogenesis genes bound by BHIH transcription factors (ChIP-Seq)
- Regulation of urea cycle genes by BZIP transcription factors (CHIP-Seq)
INTERESTED IN LEARNING HOW TO USE SPP?
Tutorials at dkNET Hypothesis Center
Watch recorded dkNET webinar on YouTube:
READ MORE